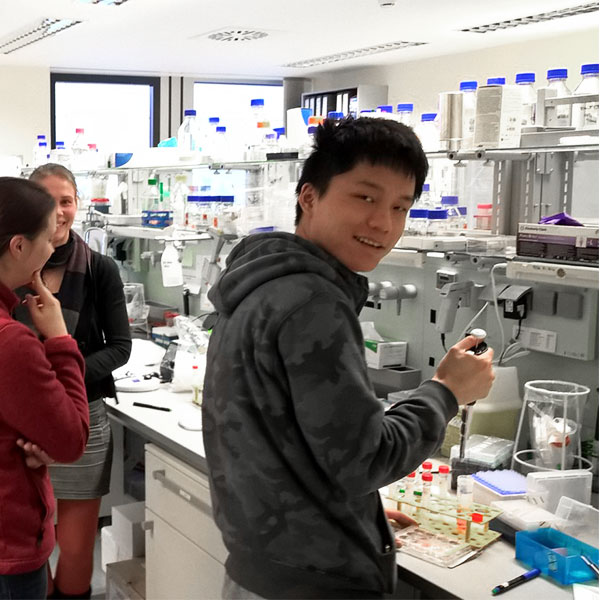
αβ T cells replacing dermal and epidermal γδ T cells in Tcrd-/- mice express an MHC-independent TCR repertoire. Binz C, Bubke A, Sandrock I, Prinz I. Eur J Immunol 2021 ; 51; 11
Case Report: Convalescent Plasma Therapy Induced Anti-SARSCoV-2 T Cell Expansion, NK Cell Maturation and Virus Clearance in a B Cell Defcient Patient After CD19 CAR T Cell Therapy. Bosnjak B, Odak I, Ritter C, Stahl K, Graalmann T, Steinbrück L, Blasczyk R, Falk CS, Schulz TF, Wedemeyer HH, Cornberg M, Ganser A, Förster R and Koenecke C Front. Immunol. 2021 Aug 12; 12:721738.
Distribution of major lymphocyte subsets and memory T-cell subpopulations in healthy adults employing GLP-conforming multicolor flow cytometry. Schultze-Florey CR, Chukhno E, Goudeva L, Blasczyk R, Ganser A, Prinz I, Förster R, Koenecke C, Odak I. Leukemia 2021; 35; 10
Clonal expansion of CD8+ T cells reflects graft-versus-leukemia activity and precedes durable remission following DLI. Schultze-Florey CR, Kuhlmann L, Raha S, Barros-Martins J, Odak I, Tan L, Xiao Y, Ravens S, Hambach L, Venturini L, Stadler M, Eder M,. Thol F, Heuser M, Förster R, Ganser A, Prinz I, Koenecke C. Blood Adv 2021 ; 5; 21
A fetal wave of human type 3 effector gd cells with restricted TCR diversity persists into adulthood. Tan L, Fichtner AS, Bruni E, Odak I, Sandrock I, Bubke A, Borchers A, Schultze-Florey C, Koenecke C, Förster R, Jarek M, von Kaisenberg C, Schulz A, Chu X, Zhang B, Li Y, Panzer U, Krebs CF, Ravens S, Prinz I. Sci. Immunol. 2021 Apr 23; 6, eabf0125 (2021).
Microbial exposure drives polyclonal expansion of innate γδ T cells immediately after birth. Ravens S, Fichtner AS, Willers M, Torkornoo D, Pirr S, Schöning J, Deseke M, Sandrock I, Bubke A, Wilharm A, Dodoo D, Egyir B, Flanagan KL, Steinbrück L, Dickinson P, Ghazal P, Adu B, Viemann D, Prinz I. Proc Natl Acad Sci U S A. 2020 Aug 4 ;117(31):18649-18660.
Reappearance of Effector 1 T Cells Is Associated With Recovery from COVID-19. Odak I, Barros-Martins J,Bošnjak B,Stahl K,David S, Wiesner O, Busch M, Hoeper MM,Pink I, Welte T, Cornberg M, Stoll M, Goudeva L, Blasczyk R, Ganser A, Prinz I, Förster R, Koenecke C*, Schultze-Florey CR.* *Authors contributed equally.EBioMedicine. 2020 Jul;57:102885.
Donor-derived IL-17A and IL-17F deficiency triggers Th1 allo-responses and increases gut leakage during acute GVHD. Odak I, Depkat-Jakob, A, Beck M, Jarek M, Yu Y, Seidler U, David S, Ganser A, Förster R, Prinz I, Koenecke C. PLoS One. 2020 Apr 6 ;15(4):e0231222.
Human γδ TCR Repertoires in Health and Disease. Fichtner AS, Ravens S, Prinz I. Cells 2020 Mar 26;9(4):E800.
Revisiting the Interaction of γδ T-Cells and B-Cells. Rampoldi, F, Ullrich L, Prinz I. Cells. 2020 Mar; 9(3): 743.
Focusing of the regulatory T-cell repertoire after allogeneic stem cell transplantation indicates protection from graft- versus-host disease. Odak I, Raha S, Schultze-Florey C, Tavil S, Ravens S, Ganser A, Förster R, Prinz I, Koenecke C. Haematologica. 2019 Dec ;104(12):e577-e580
Focusing of the regulatory T-cell repertoire after allogeneic stem cell transplantation indicates protection from graft-versus-host disease. Odak I, Raha S, Schultze-Florey C, Tavil S, Ravens S, Ganser A, Förster R, Prinz I, Koenecke C. Letters to the editor. Haematologica December 2019 104: e577-e580
Single-Cell Transcriptomics Identifies the Adaptation of Scart1+ Vγ6+ T Cells to Skin Residency as Activated Effector Cells. Tan L, Sandrock I, Odak I, Aizenbud Y, Wilharm A, Barros-Martins J, Tabib Y, Borchers A, Amado T, Gangoda L, Herold MJ, Schmidt-Supprian M, Kisielow J, Silva-Santos B, Koenecke C, Hovav AH, Krebs C, Prinz I, Ravens S. Cell Rep. 2019 Jun 18; 27(12):3657-3671.e4.
Mutual interplay between IL-17-producing γδT cells and microbiota orchestrates oral mucosal homeostasis. Wilharm A, Tabib Y, Nassar M, Reinhardt A, Mizraji G, Sandrock I, Heyman O, Barros-Martins J, Aizenbud Y, Khalaileh A, Eli-Berchoer L, Elinav E, Wilensky A, Förster R, Bercovier H, Prinz I, Hovav AH. Proc Natl Acad Sci U S A. 2019 Feb 12;116(7):2652-2661.
Genetic models reveal origin, persistence and non-redundant functions of IL-17–producing γδ T cells. Sandrock I, Reinhardt A, Ravens S, Binz C, Wilharm A, Martins J, Oberdörfer L, Tan L, Lienenklaus S, Zhang B, Naumann R, Zhuang Y, Krueger A, Förster R, Prinz I. J Exp Med. 2018 Nov 19. pii: jem.20181439.
Human γδ T Cell Receptor Repertoires in Peripheral Blood Remain Stable Despite Clearance of Persistent Hepatitis C Virus Infection by Direct-Acting Antiviral Drug Therapy. Ravens S, Hengst J, Schlapphoff V, Deterding K, Dhingra A, Schultze-Florey C, Koenecke C, Cornberg M, Wedemeyer H, Prinz I. Front Immunol. 2018 Mar 16;9:510.
Characterization of high-avidity CMV specific T-cells with differential tetramer binding co-appearing after allogeneic SCT. Ogonek J, Verma K, Schultze-Florey S, Varanasi P, Luther S, Schweier P, Kühnau W, Göhring G, Dammann E, Stadler M, Ganser A, Koehl U, Koenecke C, Weissinger EM, Hambach L. J Immunol. 2017 Jul 15; 199(2):792-805.
Eomes expression reports the progressive differentiation of IFN-γ-producing Th1-like γδ T cells. Lino CNR, Barros-Martins J, Oberdörfer L, Walzer T, Prinz I. Eur J Immunol. 2017 Jun; 47(6):970-981.
Human gd T cells are quickly reconstituted after stem-cell transplantation and show adaptive clonal expansion in response to viral infection. Ravens S, Schultze-Florey C, Raha S, Sandrock I, Drenker M, Oberdörfer L, Reinhardt A, Ravens I, Beck M, Geffers R, von Kaisenberg C, Heuser M, Thol F, Ganser A, Förster R, Koenecke C, Prinz I. Nat Immunol. 2017 Apr; 18(4):393-401.
VH1 family immunoglobulin repertoire sequencing after allogeneic hematopoietic stem cell Transplantation. Sethi MK, Thol F, Stadler M, Heuser M, Ganser A, Koenecke C and Pabst O. PLoS One. 2017 Jan 17;12(1):e0168096.
Disruption of de novo fatty acid synthesis via acetyl-CoA carboxylase 1 inhibition prevents acute graft-versus-host disease. Raha S, Raud B, Oberdörfer L, Castro CN, Schreder A, Freitag J, Longerich T, Lochner M, Sparwasser T, Berod L, Koenecke C, Prinz I. Eur J Immunol. 2016 Sep;46(9):2233-8.
Differential Effects of Gut-Homing Molecules CC Chemokine Receptor 9 and Integrin-β7 during Acute Graft-versus-Host Disease of the Liver. Schreder A, Moschovakis GL, Halle S, Schlue J, Lee CW, Schippers A, David S, Bernhardt G, Ganser A, Pabst O, Förster R, Koenecke C. Biol Blood Marrow Transplant. 2015 Dec; 21(12):2069-78.
MicroRNA-181a/b-1 Is Not Required for Innate γδ NKT Effector Cell Development. Sandrock I, Ziętara N, Łyszkiewicz M, Oberdörfer L, Witzlau K, Krueger A, Prinz I. PLoS One. 2015 Dec 16;10(12):e0145010.
γδ T cells come to stay: Innate skin memory in the Aldara model. Prinz I, Sandrock I. Eur J Immunol. 2015 Nov; 45(11):2994-7.
Impact of the revised International Prognostic Scoring System, cytogenetics and monosomal karyotype on outcome after allogeneic stem cell transplantation for myelodysplastic syndromes and secondary acute myeloid leukemia evolving from myelodysplastic syndromes: a retrospective multicenter study of the European Society of Blood and Marrow Transplantation. Koenecke C, Göhring G, de Wreede LC, van Biezen A, Scheid C, Volin L, Maertens J, Finke J, Schaap N, Robin M, Passweg J, Cornelissen J, Beelen D, Heuser M, de Witte T, Kröger N; MDS subcommittee of the Chronic Malignancies Working Party of the EBMT. Haematologica. 2015 Mar; 100(3):400-8.
A clonotypic Vγ4Jγ1/Vδ5Dδ2Jδ1 innate γδ T-cell population restricted to the CCR6⁺CD27⁻ subset. Kashani E, Föhse L, Raha S, Sandrock I, Oberdörfer L, Koenecke C, Suerbaum S, Weiss S, Prinz I. Nat Commun. 2015 Mar 13;6:6477.